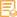Catalog: YP1840
Size
Price
Status
Qty.
200μL
$936.00
In stock
0
100μL
$560.00
In stock
0
50μL
$300.00
In stock
0
Add to cart


Collected


Collect
Main Information
Target
FANCD2 Phospho Thr691
Host Species
Rabbit
Reactivity
Human, Mouse, Rat
Applications
IHC, WB
MW
166kD (Observed)
Conjugate/Modification
Phospho
Detailed Information
Recommended Dilution Ratio
WB 1:500-2000; IHC 1:50-200
Formulation
Liquid in PBS containing 50% glycerol, and 0.02% sodium azide.
Specificity
This antibody detects endogenous levels of FANCD2 (Phospho Thr691) Rabbit pAb at Human, Mouse,Rat.The name of modified sites may be influenced by many factors, such as species (the modified site was not originally found in human samples) and the change of protein sequence (the previous protein sequence is incomplete, and the protein sequence may be prolonged with the development of protein sequencing technology). When naming, we will use the "numbers" in historical reference to keep the sites consistent with the reports. The antibody binds to the following modification sequence (lowercase letters are modification sites):YDtQD
Purification
The antibody was affinity-purified from rabbit serum by affinity-chromatography using specific immunogen.
Storage
-15°C to -25°C/1 year(Do not lower than -25°C)
Concentration
1 mg/ml
MW(Observed)
166kD
Modification
Phospho
Clonality
Polyclonal
Related Products
Antigen&Target Information
Immunogen:
Synthesized peptide derived from human FANCD2 (Phospho Thr691)
show all
Specificity:
This antibody detects endogenous levels of FANCD2 (Phospho Thr691) Rabbit pAb at Human, Mouse,Rat.The name of modified sites may be influenced by many factors, such as species (the modified site was not originally found in human samples) and the change of protein sequence (the previous protein sequence is incomplete, and the protein sequence may be prolonged with the development of protein sequencing technology). When naming, we will use the "numbers" in historical reference to keep the sites consistent with the reports. The antibody binds to the following modification sequence (lowercase letters are modification sites):YDtQD
show all
Gene Name:
FANCD2 FACD
show all
Protein Name:
Fanconi anemia group D2 protein (Protein FACD2)
show all
Other Name:
Fanconi anemia group D2 protein ;
Protein FACD2 ;
Protein FACD2 ;
show all
Database Link:
Background:
Fanconi anemia complementation group D2(FANCD2) Homo sapiens The Fanconi anemia complementation group (FANC) currently includes FANCA, FANCB, FANCC, FANCD1 (also called BRCA2), FANCD2, FANCE, FANCF, FANCG, FANCI, FANCJ (also called BRIP1), FANCL, FANCM and FANCN (also called PALB2). The previously defined group FANCH is the same as FANCA. Fanconi anemia is a genetically heterogeneous recessive disorder characterized by cytogenetic instability, hypersensitivity to DNA crosslinking agents, increased chromosomal breakage, and defective DNA repair. The members of the Fanconi anemia complementation group do not share sequence similarity; they are related by their assembly into a common nuclear protein complex. This gene encodes the protein for complementation group D2. This protein is monoubiquinated in response to DNA damage, resulting in its localization to nuclear foci with other proteins (BRCA1 AND BRCA2) involved in homology-directed DNA repai
show all
Function:
developmental stage:Highly expressed in fetal oocytes, and in hematopoietic cells of the fetal liver and bone marrow (at protein level).,Disease:Defects in FANCD2 are a cause of Fanconi anemia (FA) [MIM:227650]. FA is a genetically heterogeneous, autosomal recessive disorder characterized by progressive pancytopenia, a diverse assortment of congenital malformations, and a predisposition to the development of malignancies. At the cellular level it is associated with hypersensitivity to DNA-damaging agents, chromosomal instability (increased chromosome breakage), and defective DNA repair.,Domain:The C-terminal 24 residues of isoform 2 are required for its function.,Function:Required for maintenance of chromosomal stability. Promotes accurate and efficient pairing of homologs during meiosis. Involved in the repair of DNA double-strand breaks, both by homologous recombination and single-strand annealing. May participate in S phase and G2 phase checkpoint activation upon DNA damage. Promotes BRCA2/FANCD1 loading onto damaged chromatin. May also be involved in B-cell immunoglobulin isotype switching.,PTM:Monoubiquitinated on Lys-561 during S phase and upon genotoxic stress (isoform 1 and isoform 2). Deubiquitinated by USP1 as cells enter G2/M, or once DNA repair is completed. Monoubiquitination requires the FANCA-FANCB-FANCC-FANCE-FANCF-FANCG-FANCM complex, RPA1 and ATR, and is mediated by FANCL/PHF9. Ubiquitination is required for binding to chromatin, interaction with BRCA1 and BRCA2, DNA repair, and normal cell cycle progression, but not for phosphorylation on Ser-222 or interaction with MEN1.,PTM:Phosphorylated in response to various genotoxic stresses by ATM and/or ATR. Upon ionizing radiation, phosphorylated by ATM on Ser-222 and Ser-1404. Phosphorylation on Ser-222 is required for S-phase checkpoint activation, but not for ubiquitination, foci formation, or DNA repair. In contrast, phosphorylation by ATR on other sites may be required for ubiquitination and foci formation.,subcellular location:Concentrates in nuclear foci during S phase and upon genotoxic stress.,subunit:Interacts directly with FANCE and FANCI. Interacts with USP1 and MEN1. The ubiquitinated form specifically interacts with BRCA1, BRCA2 and BLM.,tissue specificity:Highly expressed in germinal center cells of the spleen, tonsil, and reactive lymph nodes, and in the proliferating basal layer of squamous epithelium of tonsil, esophagus, oropharynx, larynx and cervix. Expressed in cytotrophoblastic cells of the placenta and exocrine cells of the pancreas (at protein level). Highly expressed in testis, where expression is restricted to maturing spermatocytes.,
show all
Cellular Localization:
Nucleus . Concentrates in nuclear foci during S phase and upon genotoxic stress. At the onset of mitosis, excluded from chromosomes and diffuses into the cytoplasm, returning to the nucleus at the end of cell division. Observed in a few spots localized in pairs on the sister chromatids of mitotic chromosome arms and not centromeres, one on each chromatids. These foci coincide with common fragile sites and could be sites of replication fork stalling. The foci are frequently interlinked through BLM-associated ultra-fine DNA bridges. Following aphidicolin treatment, targets chromatid gaps and breaks.
show all
Research Areas:
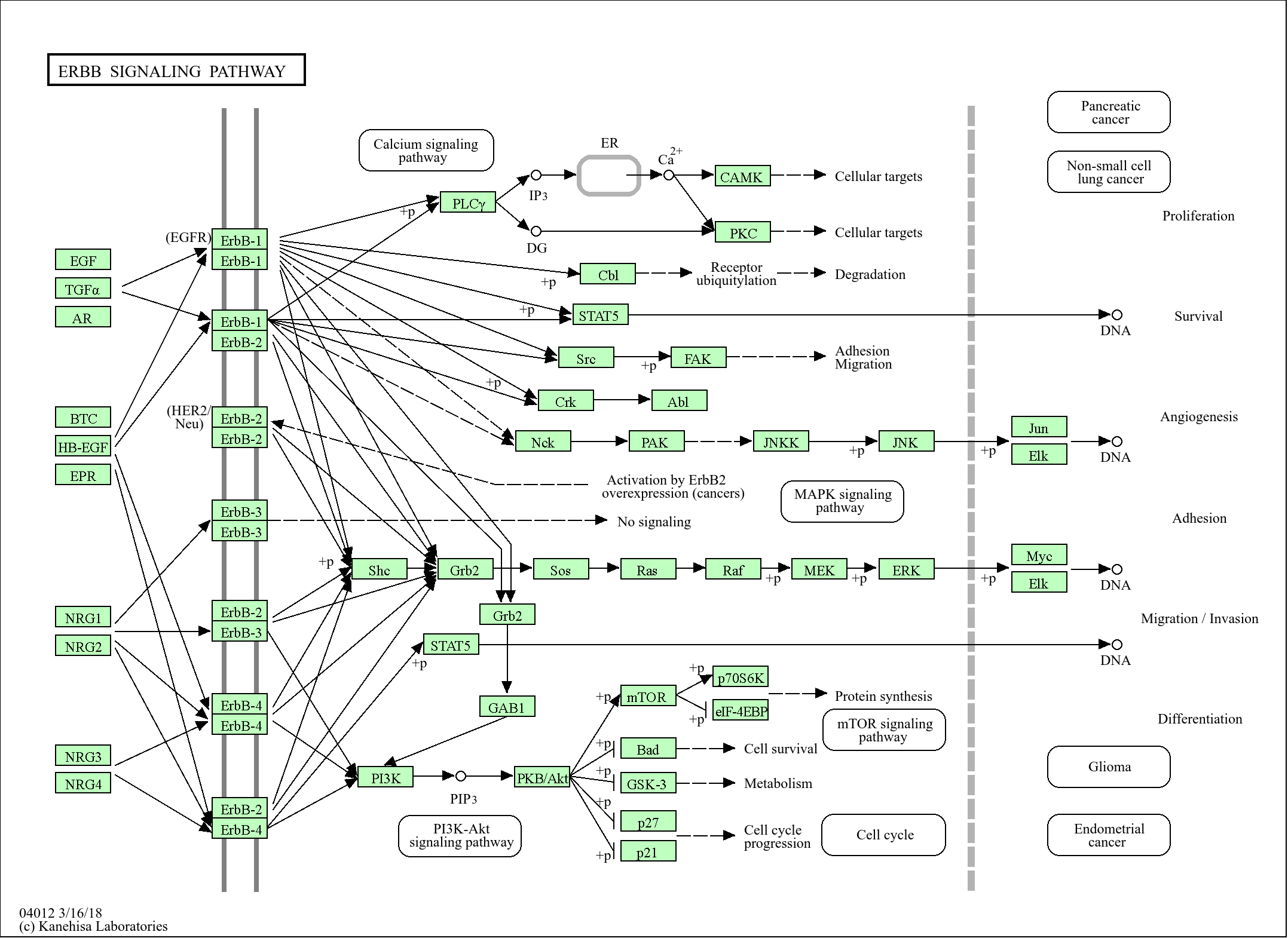
>>Fanconi anemia pathway
show all
Reference Citation({{totalcount}})
Catalog: YP1840
Size
Price
Status
Qty.
200μL
$936.00
In stock
0
100μL
$560.00
In stock
0
50μL
$300.00
In stock
0
Add to cart


Collected


Collect
Recently Viewed Products
Clear allPRODUCTS
CUSTOMIZED
ABOUT US
Toggle night Mode
{{pinfoXq.title || ''}}
Catalog: {{pinfoXq.catalog || ''}}
Filter:
All
{{item.name}}
{{pinfo.title}}
-{{pinfo.catalog}}
Main Information
Target
{{pinfo.target}}
Reactivity
{{pinfo.react}}
Applications
{{pinfo.applicat}}
Conjugate/Modification
{{pinfo.coupling}}/{{pinfo.modific}}
MW (kDa)
{{pinfo.mwcalc}}
Host Species
{{pinfo.hostspec}}
Isotype
{{pinfo.isotype}}
Product {{index}}/{{pcount}}
Prev
Next
{{pvTitle}}
Scroll wheel zooms the picture
{{pvDescr}}