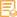Catalog: YP1383
Size
Price
Status
Qty.
200μL
$600.00
3 weeks
0
100μL
$340.00
3 weeks
0
50μL
$190.00
3 weeks
0
Add to cart


Collected


Collect
Main Information
Target
LATS1 Phospho Thr1079
Host Species
Rabbit
Reactivity
Human, Mouse
Applications
WB, ELISA, IHC
MW
140kD (Observed)
Conjugate/Modification
Phospho
Detailed Information
Recommended Dilution Ratio
WB 1:500-2000; IHC 1:50-300; ELISA 1:2000-20000
Formulation
Liquid in PBS containing 50% glycerol, 0.5% BSA and 0.02% sodium azide.
Specificity
This antibody detects endogenous levels of Human Mouse LATS1 (phospho-Thr1079)
Purification
The antibody was affinity-purified from rabbit serum by affinity-chromatography using specific immunogen.
Storage
-15°C to -25°C/1 year(Do not lower than -25°C)
Concentration
1 mg/ml
MW(Observed)
140kD
Modification
Phospho
Clonality
Polyclonal
Isotype
IgG
Related Products
Antigen&Target Information
Immunogen:
Synthesized phosho peptide around human LATS1 (Thr1079)
show all
Specificity:
This antibody detects endogenous levels of Human Mouse LATS1 (phospho-Thr1079)
show all
Gene Name:
LATS1 WARTS
show all
Protein Name:
LATS1 (Thr1079)
show all
Other Name:
Serine/threonine-protein kinase LATS1 ;
Large tumor suppressor homolog 1 ;
WARTS protein kinase ;
h-warts ;
Large tumor suppressor homolog 1 ;
WARTS protein kinase ;
h-warts ;
show all
Background:
The protein encoded by this gene is a putative serine/threonine kinase that localizes to the mitotic apparatus and complexes with cell cycle controller CDC2 kinase in early mitosis. The protein is phosphorylated in a cell-cycle dependent manner, with late prophase phosphorylation remaining through metaphase. The N-terminal region of the protein binds CDC2 to form a complex showing reduced H1 histone kinase activity, indicating a role as a negative regulator of CDC2/cyclin A. In addition, the C-terminal kinase domain binds to its own N-terminal region, suggesting potential negative regulation through interference with complex formation via intramolecular binding. Biochemical and genetic data suggest a role as a tumor suppressor. This is supported by studies in knockout mice showing development of soft-tissue sarcomas, ovarian stromal cell tumors and a high sensitivity to carcinogenic treatmen
show all
Function:
Catalytic activity:ATP + a protein = ADP + a phosphoprotein.,cofactor:Magnesium.,Function:Tumor suppressor which plays a critical role in maintenance of ploidy through its actions in both mitotic progression and the G1 tetraploidy checkpoint. Negatively regulates G2/M transition by down-regulating CDC2 kinase activity. Involved in the control of p53 expression. Affects cytokinesis by regulating actin polymerization through negative modulation of LIMK1. May also play a role in endocrine function.,PTM:Autophosphorylated and phosphorylated during M-phase of the cell cycle. Phosphorylated by STK3 at Ser-909 and Thr-1079, which results in its activation. Phosphorylated upon DNA damage, probably by ATM or ATR.,similarity:Belongs to the protein kinase superfamily. AGC Ser/Thr protein kinase family.,similarity:Contains 1 AGC-kinase C-terminal domain.,similarity:Contains 1 protein kinase domain.,similarity:Contains 1 UBA domain.,subcellular location:Localizes to the centrosomes throughout interphase but migrates to the mitotic apparatus, including spindle pole bodies, mitotic spindle, and midbody, during mitosis.,subunit:Complexes with CDC2 in early mitosis. LATS1-associated CDC2 has no mitotic cyclin partner and no apparent kinase activity. Binds phosphorylated ZYX, locating this protein to the mitotic spindle and suggesting a role for actin regulatory proteins during mitosis. Binds to and colocalizes with LIMK1 at the actomyosin contractile ring during cytokinesis.,tissue specificity:Expressed in all adult tissues examined except for lung and kidney.,
show all
Cellular Localization:
Cytoplasm, cytoskeleton, microtubule organizing center, centrosome . Cytoplasm, cytoskeleton, spindle . Midbody . Cytoplasm, cytoskeleton, microtubule organizing center, spindle pole body . Localizes to the centrosomes throughout interphase but migrates to the mitotic apparatus, including spindle pole bodies, mitotic spindle, and midbody, during mitosis. .
show all
Tissue Expression:
Research Areas:
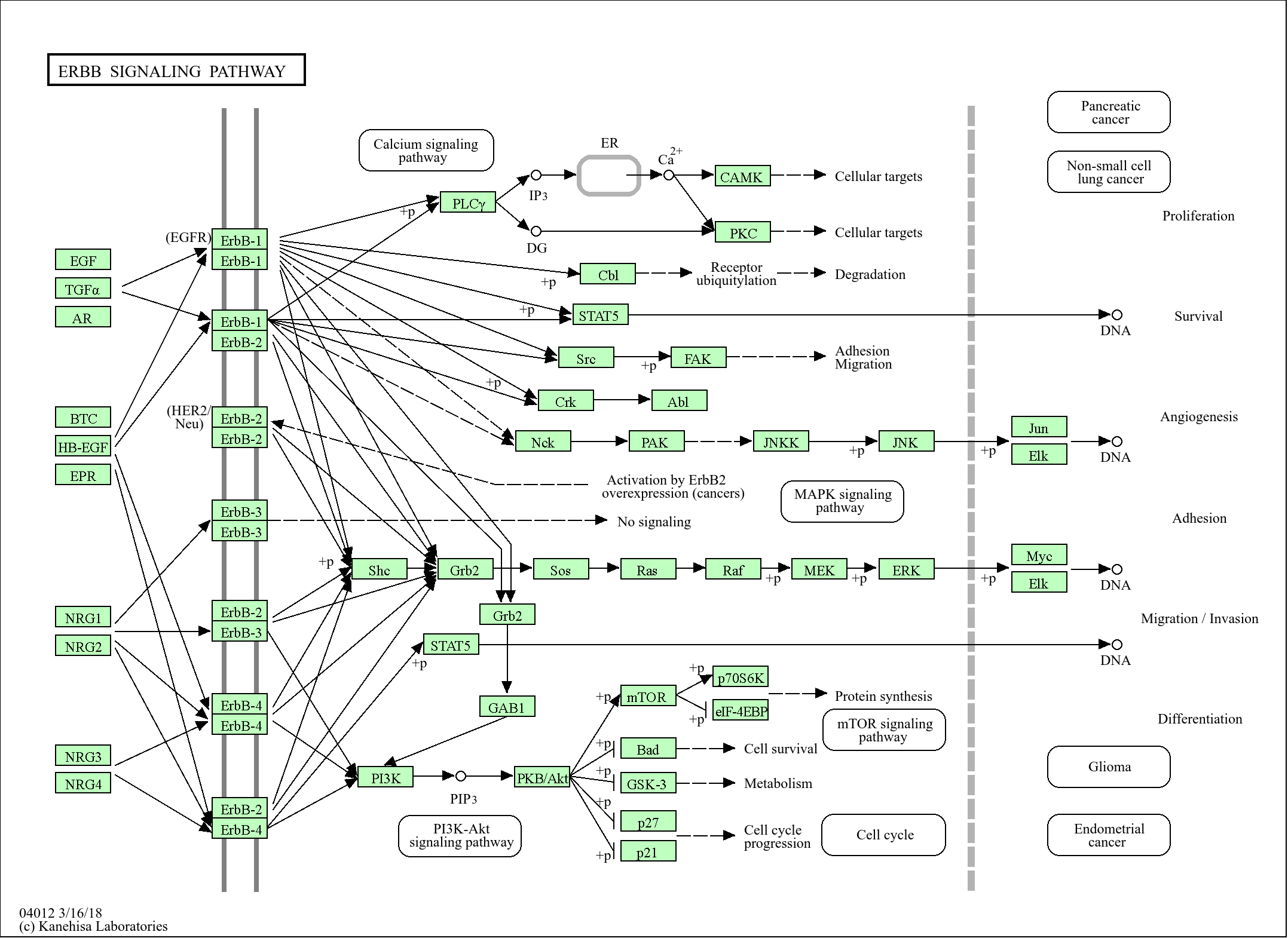
>>Hippo signaling pathway ;
>>Hippo signaling pathway - multiple species
>>Hippo signaling pathway - multiple species
show all
Signaling Pathway
Reference Citation({{totalcount}})
Catalog: YP1383
Size
Price
Status
Qty.
200μL
$600.00
3 weeks
0
100μL
$340.00
3 weeks
0
50μL
$190.00
3 weeks
0
Add to cart


Collected


Collect
Recently Viewed Products
Clear allPRODUCTS
CUSTOMIZED
ABOUT US
Toggle night Mode
{{pinfoXq.title || ''}}
Catalog: {{pinfoXq.catalog || ''}}
Filter:
All
{{item.name}}
{{pinfo.title}}
-{{pinfo.catalog}}
Main Information
Target
{{pinfo.target}}
Reactivity
{{pinfo.react}}
Applications
{{pinfo.applicat}}
Conjugate/Modification
{{pinfo.coupling}}/{{pinfo.modific}}
MW (kDa)
{{pinfo.mwcalc}}
Host Species
{{pinfo.hostspec}}
Isotype
{{pinfo.isotype}}
Product {{index}}/{{pcount}}
Prev
Next
{{pvTitle}}
Scroll wheel zooms the picture
{{pvDescr}}