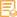Catalog: YP0121
Size
Price
Status
Qty.
200μL
$600.00
In stock
0
100μL
$340.00
In stock
0
50μL
$190.00
In stock
0
Add to cart


Collected


Collect
Main Information
Target
GRF-1 Phospho Tyr1087
Host Species
Rabbit
Reactivity
Human, Mouse, Rat
Applications
WB, ELISA
MW
190kD (Observed)
Conjugate/Modification
Phospho
Detailed Information
Recommended Dilution Ratio
WB 1:500-1:2000; ELISA 1:10000; Not yet tested in other applications.
Formulation
Liquid in PBS containing 50% glycerol, 0.5% BSA and 0.02% sodium azide.
Specificity
Phospho-GRF-1 (Y1087) Polyclonal Antibody detects endogenous levels of GRF-1 protein only when phosphorylated at Y1087.The name of modified sites may be influenced by many factors, such as species (the modified site was not originally found in human samples) and the change of protein sequence (the previous protein sequence is incomplete, and the protein sequence may be prolonged with the development of protein sequencing technology). When naming, we will use the "numbers" in historical reference to keep the sites consistent with the reports. The antibody binds to the following modification sequence (lowercase letters are modification sites):SDyAE
Purification
The antibody was affinity-purified from rabbit antiserum by affinity-chromatography using epitope-specific immunogen.
Storage
-15°C to -25°C/1 year(Do not lower than -25°C)
Concentration
1 mg/ml
MW(Observed)
190kD
Modification
Phospho
Clonality
Polyclonal
Isotype
IgG
Related Products
Antigen&Target Information
Immunogen:
Synthesized phospho-peptide around the phosphorylation site of human GRF-1 (phospho Tyr1087)
show all
Specificity:
Phospho-GRF-1 (Y1087) Polyclonal Antibody detects endogenous levels of GRF-1 protein only when phosphorylated at Y1087.The name of modified sites may be influenced by many factors, such as species (the modified site was not originally found in human samples) and the change of protein sequence (the previous protein sequence is incomplete, and the protein sequence may be prolonged with the development of protein sequencing technology). When naming, we will use the "numbers" in historical reference to keep the sites consistent with the reports. The antibody binds to the following modification sequence (lowercase letters are modification sites):SDyAE
show all
Gene Name:
ARHGAP35
show all
Protein Name:
Rho GTPase-activating protein 35
show all
Other Name:
ARHGAP35 ;
GRF1 ;
GRLF1 ;
KIAA1722 ;
Rho GTPase-activating protein 35 ;
Glucocorticoid receptor DNA-binding factor 1 ;
Glucocorticoid receptor repression factor 1 ;
GRF-1 ;
Rho GAP p190A ;
p190-A
GRF1 ;
GRLF1 ;
KIAA1722 ;
Rho GTPase-activating protein 35 ;
Glucocorticoid receptor DNA-binding factor 1 ;
Glucocorticoid receptor repression factor 1 ;
GRF-1 ;
Rho GAP p190A ;
p190-A
show all
Background:
The human glucocorticoid receptor DNA binding factor, which associates with the promoter region of the glucocorticoid receptor gene (hGR gene), is a repressor of glucocorticoid receptor transcription. The amino acid sequence deduced from the cDNA sequences show the presence of three sequence motifs characteristic of a zinc finger and one motif suggestive of a leucine zipper in which 1 cysteine is found instead of all leucines. The GRLF1 enhances the homologous down-regulation of wild-type hGR gene expression. Biochemical analysis suggests that GRLF1 interaction is sequence specific and that transcriptional efficacy of GRLF1 is regulated through its interaction with specific sequence motif. The level of expression is regulated by glucocorticoids. [provided by RefSeq, Jul 2008],
show all
Function:
Function:Represses transcription of the glucocorticoid receptor by binding to the cis-acting regulatory sequence 5'-GAGAAAAGAAACTGGAGAAACTC-3'. May participate in the regulation of retinal development and degeneration. May transduce signals from p21-ras to the nucleus, acting via the ras GTPase-activating protein (GAP). May also act as a tumor suppressor.,PTM:Phosphorylated upon DNA damage, probably by ATM or ATR.,PTM:Tyrosine phosphorylated.,similarity:Contains 1 Rho-GAP domain.,similarity:Contains 4 FF domains.,subunit:Interacts with p120GAP.,
show all
Cellular Localization:
Cytoplasm, cytoskeleton, cilium basal body . Cytoplasm . Nucleus . Cell membrane . In response to integrins and SDC4 and upon phosphorylation by PKC, relocalizes from the cytoplasm to regions of plasma membrane ruffling where it colocalizes with polymerized actin. .
show all
Tissue Expression:
Research Areas:
>>Focal adhesion ;
>>Platelet activation ;
>>Leukocyte transendothelial migration ;
>>Regulation of actin cytoskeleton
>>Platelet activation ;
>>Leukocyte transendothelial migration ;
>>Regulation of actin cytoskeleton
show all
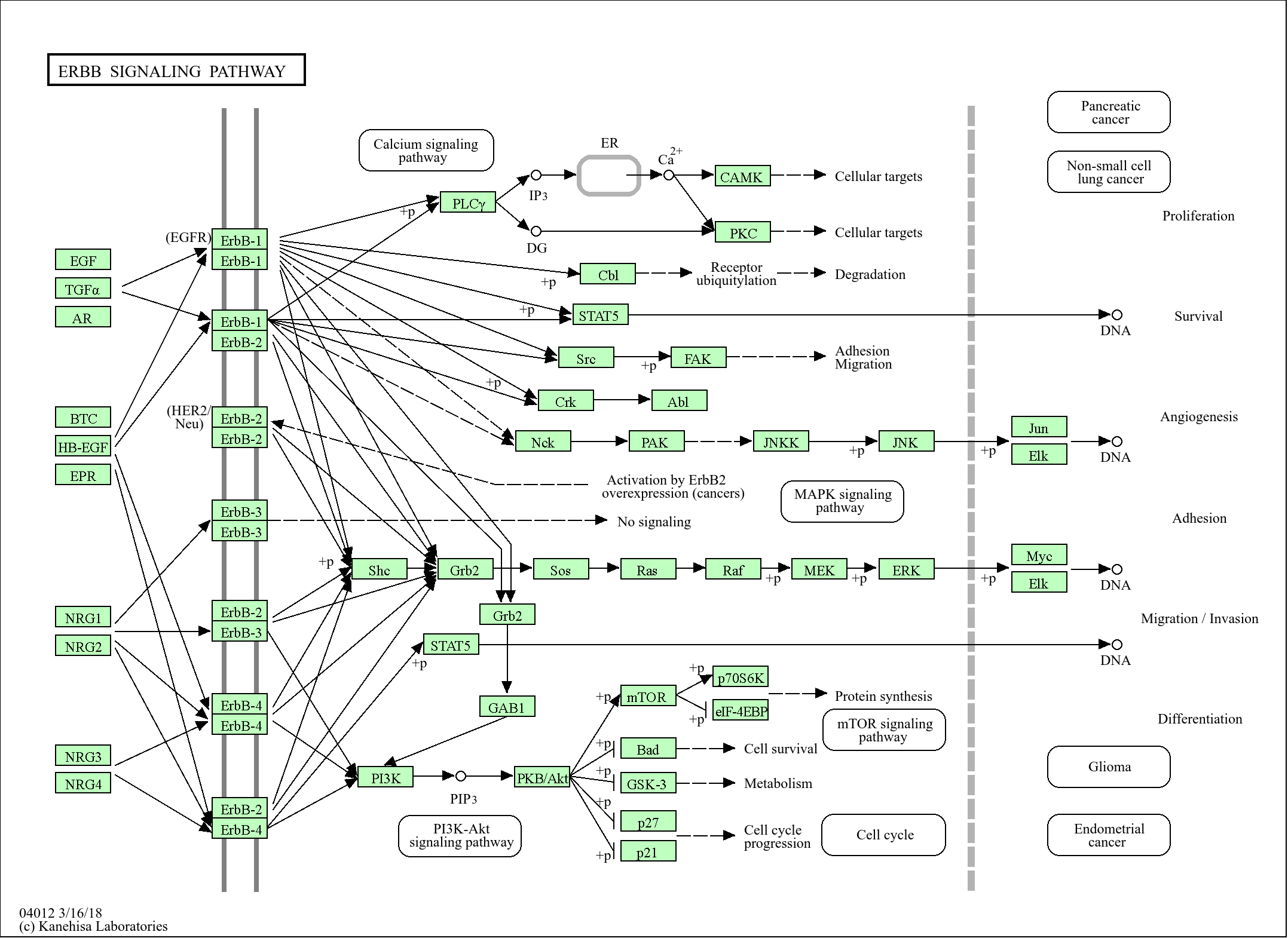
Signaling Pathway
Reference Citation({{totalcount}})
Catalog: YP0121
Size
Price
Status
Qty.
200μL
$600.00
In stock
0
100μL
$340.00
In stock
0
50μL
$190.00
In stock
0
Add to cart


Collected


Collect
Recently Viewed Products
Clear allPRODUCTS
CUSTOMIZED
ABOUT US
Toggle night Mode
{{pinfoXq.title || ''}}
Catalog: {{pinfoXq.catalog || ''}}
Filter:
All
{{item.name}}
{{pinfo.title}}
-{{pinfo.catalog}}
Main Information
Target
{{pinfo.target}}
Reactivity
{{pinfo.react}}
Applications
{{pinfo.applicat}}
Conjugate/Modification
{{pinfo.coupling}}/{{pinfo.modific}}
MW (kDa)
{{pinfo.mwcalc}}
Host Species
{{pinfo.hostspec}}
Isotype
{{pinfo.isotype}}
Product {{index}}/{{pcount}}
Prev
Next
{{pvTitle}}
Scroll wheel zooms the picture
{{pvDescr}}