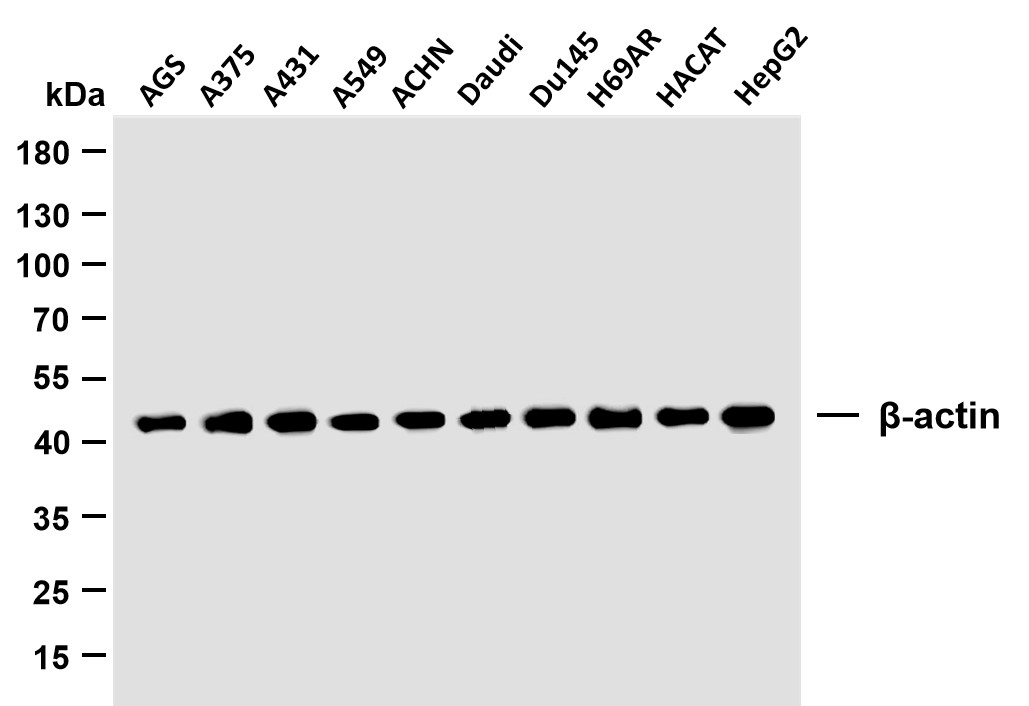
Centaurin-β1 rabbit pAb
- Catalog No.:YT7777
- Applications:WB;ELISA
- Reactivity:Human;Rat;Mouse;
- Target:
- Centaurin-β1
- Fields:
- >>Endocytosis
- Gene Name:
- ACAP1 CENTB1 KIAA0050
- Protein Name:
- Centaurin-β1
- Human Gene Id:
- 9744
- Human Swiss Prot No:
- Q15027
- Mouse Gene Id:
- 216859
- Mouse Swiss Prot No:
- Q8K2H4
- Immunogen:
- Synthesized peptide derived from human Centaurin-β1 AA range: 490-570
- Specificity:
- This antibody detects endogenous levels of Human Centaurin-β1
- Formulation:
- Liquid in PBS containing 50% glycerol, 0.5% BSA and 0.02% sodium azide.
- Source:
- Polyclonal, Rabbit,IgG
- Dilution:
- WB 1:1000-2000 ELISA 1:5000-20000
- Purification:
- The antibody was affinity-purified from rabbit antiserum by affinity-chromatography using epitope-specific immunogen.
- Concentration:
- 1 mg/ml
- Storage Stability:
- -15°C to -25°C/1 year(Do not lower than -25°C)
- Other Name:
- Arf-GAP with coiled-coil, ANK repeat and PH domain-containing protein 1 (Centaurin-beta-1;Cnt-b1)
- Molecular Weight(Da):
- 81kD
- Background:
- domain:PH domain binds phospholipids including phosphatidic acid, phosphatidylinositol 3-phosphate, phosphatidylinositol 3,5-bisphosphate (PIP2) and phosphatidylinositol 3,4,5-trisphosphate (PIP3). May mediate ACAP1-binding to PIP2 or PIP3 containing membranes.,enzyme regulation:GAP activity stimulated by phosphatidylinositol 4,5-bisphosphate (PIP2) and phosphatidic acid.,function:GTPase-activating protein (GAP) for ADP ribosylation factor 6 (ARF6) required for clathrin-dependent export of proteins from recycling endosomes to trans-Golgi network and cell surface.,miscellaneous:Cells overexpressing ACAP1 show an accumulation of ITGB1 in recycling endosomes and inhibition of stimulation-dependent cell migration. Cells with reduced levels of ACAP1 or AKT1 and AKT2 show inhibition of stimulation-dependent cell migration. Cells overexpressing ACAP1 and PIP5K1C show formation of tubular structures derived from endosomal membranes.,PTM:Phosphorylation at Ser-554 by PKB is required for interaction with ITGB1, export of ITGB1 from recycling endosomes to the cell surface and ITGB1-dependent cell migration.,similarity:Contains 1 Arf-GAP domain.,similarity:Contains 1 BAR domain.,similarity:Contains 1 PH domain.,similarity:Contains 3 ANK repeats.,subunit:Interacts with GTP-bound ARF6. Interacts with third cytoplasmic loop of SLC2A4/GLUT4. Interacts with CLTC. Interacts with GULP1. Forms a complex with GDP-bound ARF6 and GULP1.,tissue specificity:Highest level in lung and spleen. Low level in heart, kidney, liver and pancreas.,
- Function:
- domain:PH domain binds phospholipids including phosphatidic acid, phosphatidylinositol 3-phosphate, phosphatidylinositol 3,5-bisphosphate (PIP2) and phosphatidylinositol 3,4,5-trisphosphate (PIP3). May mediate ACAP1-binding to PIP2 or PIP3 containing membranes.,enzyme regulation:GAP activity stimulated by phosphatidylinositol 4,5-bisphosphate (PIP2) and phosphatidic acid.,function:GTPase-activating protein (GAP) for ADP ribosylation factor 6 (ARF6) required for clathrin-dependent export of proteins from recycling endosomes to trans-Golgi network and cell surface.,miscellaneous:Cells overexpressing ACAP1 show an accumulation of ITGB1 in recycling endosomes and inhibition of stimulation-dependent cell migration. Cells with reduced levels of ACAP1 or AKT1 and AKT2 show inhibition of stimulation-dependent cell migration. Cells overexpressing ACAP1 and PIP5K1C show formation of tubular struct
- Subcellular Location:
- Recycling endosome membrane ; Peripheral membrane protein ; Cytoplasmic side .
- Expression:
- Highest level in lung and spleen. Low level in heart, kidney, liver and pancreas.
- June 19-2018
- WESTERN IMMUNOBLOTTING PROTOCOL
- June 19-2018
- IMMUNOHISTOCHEMISTRY-PARAFFIN PROTOCOL
- June 19-2018
- IMMUNOFLUORESCENCE PROTOCOL
- September 08-2020
- FLOW-CYTOMEYRT-PROTOCOL
- May 20-2022
- Cell-Based ELISA│解您多样本WB检测之困扰
- July 13-2018
- CELL-BASED-ELISA-PROTOCOL-FOR-ACETYL-PROTEIN
- July 13-2018
- CELL-BASED-ELISA-PROTOCOL-FOR-PHOSPHO-PROTEIN
- July 13-2018
- Antibody-FAQs