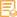Catalog: YM9239
Size
Price
Status
Qty.
200μL
$600.00
3 weeks
0
100μL
$340.00
3 weeks
0
40μL
$190.00
3 weeks
0
Add to cart


Collected


Collect
Main Information
Target
PABP1
Host Species
Rabbit
Reactivity
Human, Mouse, Rat
Applications
WB, IHC, IF, IP, ELISA
MW
71kD (Calculated)
71kD (Observed)
Conjugate/Modification
Unmodified
Detailed Information
Recommended Dilution Ratio
IHC 1:200-1:1000; WB 1:2000-1:10000; IF 1:200-1:1000; ELISA 1:5000-1:20000; IP 1:50-1:200;
Formulation
PBS, 50% glycerol, 0.05% Proclin 300, 0.05%BSA
Specificity
Endogenous
Purification
Protein A
Storage
-15°C to -25°C/1 year(Do not lower than -25°C)
MW(Calculated)
71kD
MW(Observed)
71kD
Modification
Unmodified
Clonality
Monoclonal
Clone Number
PT1397R
Isotype
IgG,Kappa
Related Products
Antigen&Target Information
Specificity:
Endogenous
show all
Gene Name:
PABPC1 PAB1 PABP1 PABPC2
show all
Protein Name:
Polyadenylate-binding protein 1 (PABP-1) (Poly(A)-binding protein 1)
show all
Background:
This gene encodes a poly(A) binding protein. The protein shuttles between the nucleus and cytoplasm and binds to the 3' poly(A) tail of eukaryotic messenger RNAs via RNA-recognition motifs. The binding of this protein to poly(A) promotes ribosome recruitment and translation initiation; it is also required for poly(A) shortening which is the first step in mRNA decay. The gene is part of a small gene family including three protein-coding genes and several pseudogenes.[provided by RefSeq, Aug 2010],
show all
Function:
Caution:Was originally (Ref.4) termed polyadenylate binding protein II.,Domain:The RNA-binding domains RRM1 and RRM2 and the C-terminus (last 138 amino acids) regions interact with the PABPC1-interacting motif-1 (PAM1) and -2 (PAM2) of PAIP1, respectively.,Domain:The RNA-binding domains RRM2 and RRM3 and the C-terminus (last 138 amino acids) regions interact with the PABPC1-interacting motif-1 (PAM1) and -2 (PAM2) of PAIP2, respectively.,Function:Binds the poly(A) tail of mRNA. May be involved in cytoplasmic regulatory processes of mRNA metabolism such as pre-mRNA splicing. Its function in translational initiation regulation can either be enhanced by PAIP1 or repressed by PAIP2. Can probably bind to cytoplasmic RNA sequences other than poly(A) in vivo. May be involved in translationally coupled mRNA turnover. Implicated with other RNA-binding proteins in the cytoplasmic deadenylation/translational and decay interplay of the FOS mRNA mediated by the major coding-region determinant of instability (mCRD) domain.,PTM:Arg-493 is dimethylated, probably to asymmetric dimethylarginine.,PTM:Methylated by CARM1.,similarity:Belongs to the polyadenylate-binding protein type-1 family.,similarity:Contains 1 PABC domain.,similarity:Contains 4 RRM (RNA recognition motif) domains.,subcellular location:Shuttles between the cytoplasm and the nucleus. Relocalizes to the nucleus upon rotavirus A infection.,subunit:Component of a multi subunit autoregulatory ribonucleoprotein complex (ARC), at least composed of IGF2BP1, PABPC1 and CSDE1. Interacts with IGF2BP1. Part of a complex associated with the FOS mCRD domain and consisting of HNRPD, SYNCRIP, PAIP1 and CSDE1/UNR. Interacts with the PABPC1-interacting motif-1 (PAM1) and -2 (PAM2) of PAIP1 and PAIP2. Interacts with PAIP1 with a 1:1 stoichiometry and with PAIP2 with a 1:2 stoichiometry. The interaction with CSDE1 is direct and RNA-independent. Found in a mRNP complex with YBX2. Interacts with PAPD4/GLD2. Identified in the spliceosome C complex, at least composed of AQR, ASCC3L1, C19orf29, CDC40, CDC5L, CRNKL1, DDX23, DDX41, DDX48, DDX5, DGCR14, DHX35, DHX38, DHX8, EFTUD2, FRG1, GPATC1, HNRPA1, HNRPA2B1, HNRPA3, HNRPC, HNRPF, HNRPH1, HNRPK, HNRPM, HNRPR, HNRPU, KIAA1160, KIAA1604, LSM2, LSM3, MAGOH, MORG1, PABPC1, PLRG1, PNN, PPIE, PPIL1, PPIL3, PPWD1, PRPF19, PRPF4B, PRPF6, PRPF8, RALY, RBM22, RBM8A, RBMX, SART1, SF3A1, SF3A2, SF3A3, SF3B1, SF3B2, SF3B3, SFRS1, SKIV2L2, SNRPA1, SNRPB, SNRPB2, SNRPD1, SNRPD2, SNRPD3, SNRPE, SNRPF, SNRPG, SNW1, SRRM1, SRRM2, SYF2, SYNCRIP, TFIP11, THOC4, U2AF1, WDR57, XAB2 and ZCCHC8. Interacts with PIWIL1 (By similarity). Interacts with NFX1.,tissue specificity:Ubiquitous.,
show all
Cellular Localization:
Cytoplasm . Cytoplasm, Stress granule . Nucleus . Cell projection, lamellipodium . Localized in cytoplasmic mRNP granules containing untranslated mRNAs (PubMed:17289661). Shuttles between the cytoplasm and the nucleus (PubMed:9582337). During stress and in the absence of DDX3X, localizes to the nucleus (PubMed:21883093). At the leading edge of migrating fibroblasts, colocalizes with DDX3X (PubMed:28733330). Relocalizes to cytoplasmic stress granules upon cellular stress where it colocalizes with ENDOV (PubMed:27573237). In case of HRSV infection, localizes in cytoplasmic inclusion bodies substructures called inclusion bodies associated granules (IBAGs) (PubMed:31649314). .
show all
Tissue Expression:
show all
Research Areas:
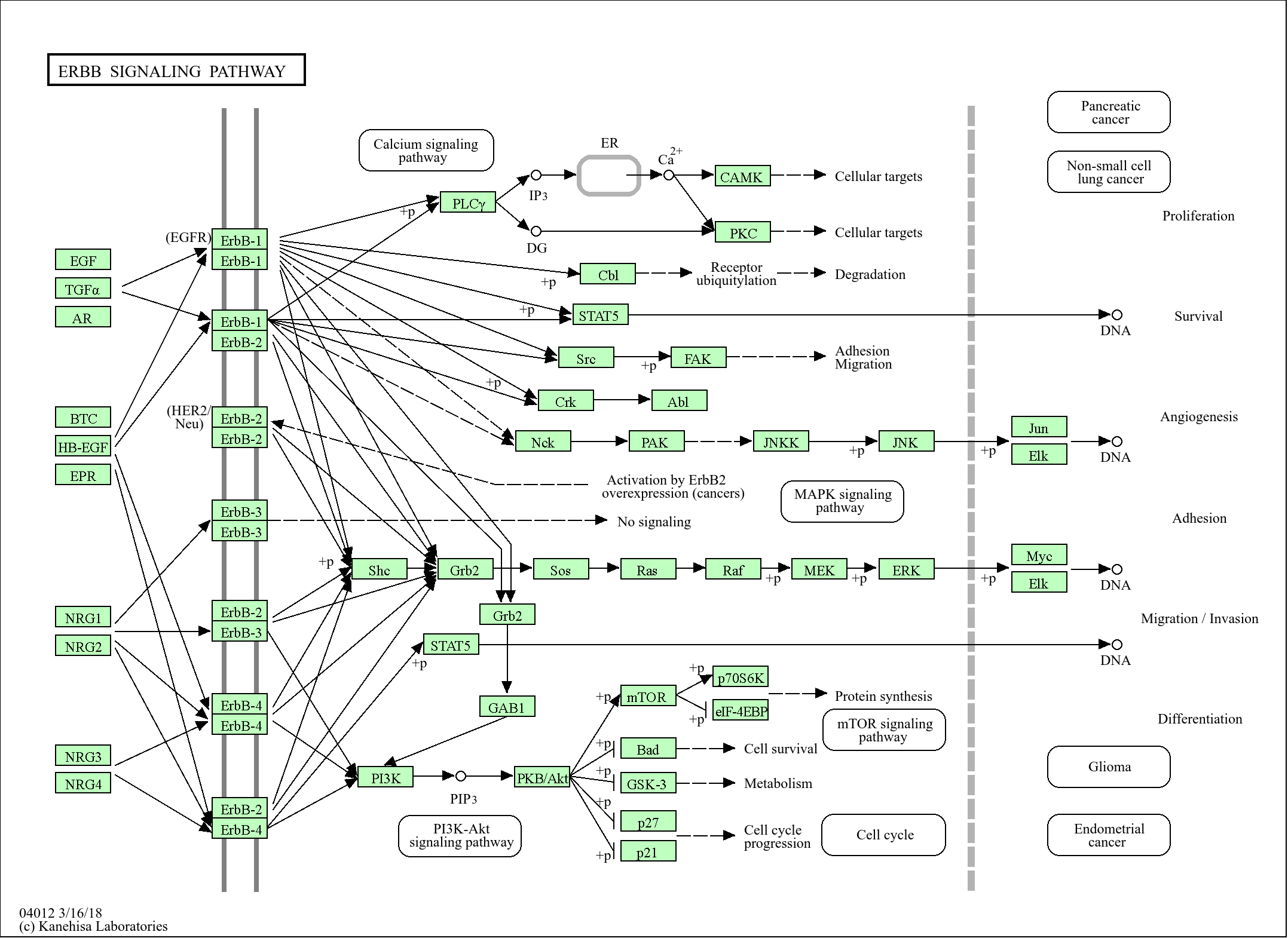
>>mRNA surveillance pathway ;
>>RNA degradation
>>RNA degradation
show all
Signaling Pathway
Reference Citation({{totalcount}})
Catalog: YM9239
Size
Price
Status
Qty.
200μL
$600.00
3 weeks
0
100μL
$340.00
3 weeks
0
40μL
$190.00
3 weeks
0
Add to cart


Collected


Collect
Recently Viewed Products
Clear allPRODUCTS
CUSTOMIZED
ABOUT US
Toggle night Mode
{{pinfoXq.title || ''}}
Catalog: {{pinfoXq.catalog || ''}}
Filter:
All
{{item.name}}
{{pinfo.title}}
-{{pinfo.catalog}}
Main Information
Target
{{pinfo.target}}
Reactivity
{{pinfo.react}}
Applications
{{pinfo.applicat}}
Conjugate/Modification
{{pinfo.coupling}}/{{pinfo.modific}}
MW (kDa)
{{pinfo.mwcalc}}
Host Species
{{pinfo.hostspec}}
Isotype
{{pinfo.isotype}}
Product {{index}}/{{pcount}}
Prev
Next
{{pvTitle}}
Scroll wheel zooms the picture
{{pvDescr}}